Browser
Protein data
function: translocaseexperiment: ELECTRON MICROSCOPY
resolution: 2.52 Å
origomeric count: 8
axial ligand #1: HIS
chainID: a,resSeq: 186,
Coordination distance[Å]: 1.949
molecule: Cytochrome bd-I ubiquinol oxidase subunit 1
EC number: 7.1.1.7
Organism: Escherichia coli K-12
axial ligand #2: MET
chainID: a,resSeq: 393,
Coordination distance[Å]: 2.799
molecule: same as ligand #1
List of other hemes in pdb:9sfj
| ID of heme | Distortion | Axial ligands on heme | Function & structure | |
|---|---|---|---|---|
9sfj-A-601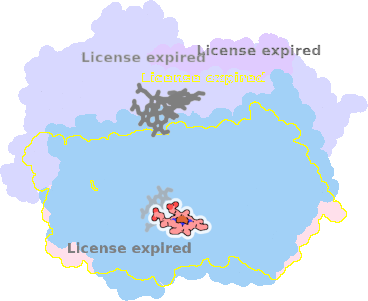 |
sad. -0.39 ruf. +0.51 dom. -0.21 bre. -0.16 |
HIS | chainID: A, resSeq: 186, molecule: Cytochrome bd-I ubiquinol oxidase subunit 1 |
translocase oligomeric count: 8 pocket vol.: 466.0 Å3 d(Fe-oop): 0.158 Å |
| MET | chainID: A, resSeq: 393, molecule: Cytochrome bd-I ubiquinol oxidase subunit 1 |
|||
9sfj-A-604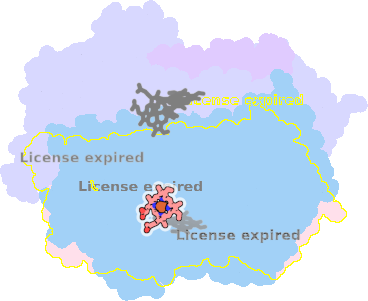 |
sad. +1.00 ruf. +0.09 dom. -0.51 bre. -0.08 |
GLU | chainID: A, resSeq: 445, molecule: Cytochrome bd-I ubiquinol oxidase subunit 1 |
translocase oligomeric count: 8 pocket vol.: 433.0 Å3 d(Fe-oop): 0.093 Å |
| Not assigned. | ||||
9sfj-a-603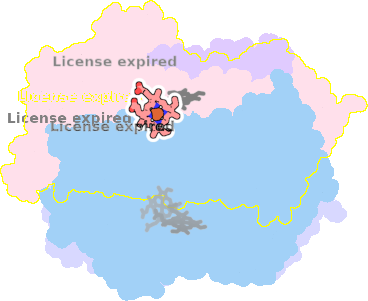 |
sad. +1.03 ruf. +0.17 dom. -0.41 bre. -0.09 |
GLU | chainID: a, resSeq: 445, molecule: Cytochrome bd-I ubiquinol oxidase subunit 1 |
translocase oligomeric count: 8 pocket vol.: 438.0 Å3 d(Fe-oop): 0.003 Å |
| Not assigned. | ||||
