Browser
Protein data
function: oxidoreductaseexperiment: X-RAY DIFFRACTION
resolution: 1.9 Å
origomeric count: 6
axial ligand #1: HIS
chainID: D,resSeq: 225,
Coordination distance[Å]: 2.093
molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup
Organism: Streptomyces lividans 1326
axial ligand #2: O
chainID: D,resSeq: 402,
Coordination distance[Å]: 1.816
molecule: OXYGEN ATOM
List of other hemes in pdb:9qqy
| ID of heme | Distortion | Axial ligands on heme | Function & structure | |
|---|---|---|---|---|
9qqy-A-401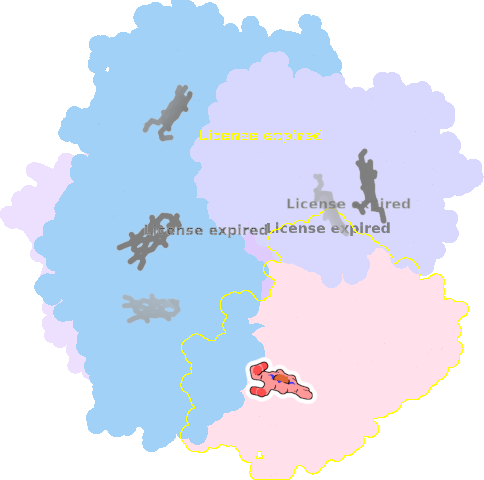 |
sad. +0.28 ruf. +0.32 dom. +0.11 bre. -0.09 |
HIS | chainID: A, resSeq: 225, molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup |
oxidoreductase oligomeric count: 6 pocket vol.: 414.0 Å3 d(Fe-oop): 0.035 Å |
| O | chainID: A, resSeq: 403, molecule: OXYGEN ATOM |
|||
9qqy-B-401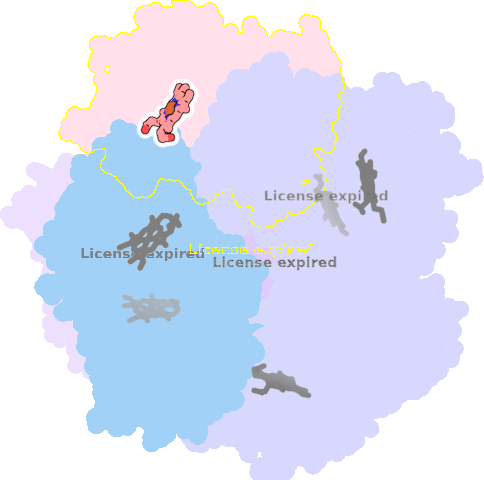 |
sad. +0.21 ruf. +0.19 dom. +0.05 bre. -0.23 |
HIS | chainID: B, resSeq: 225, molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup |
oxidoreductase oligomeric count: 6 pocket vol.: 422.0 Å3 d(Fe-oop): 0.044 Å |
| O | chainID: B, resSeq: 402, molecule: OXYGEN ATOM |
|||
9qqy-C-401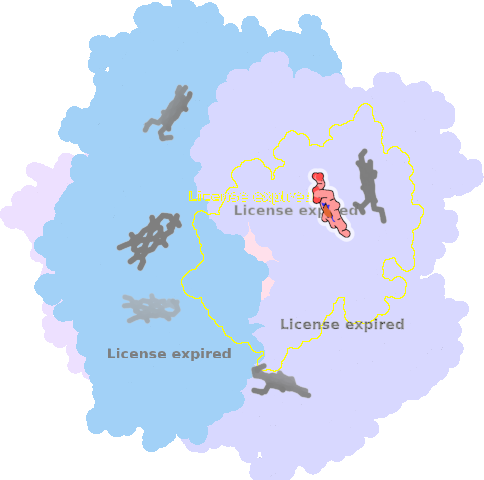 |
sad. +0.32 ruf. +0.34 dom. +0.08 bre. -0.09 |
HIS | chainID: C, resSeq: 225, molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup |
oxidoreductase oligomeric count: 6 pocket vol.: 436.0 Å3 d(Fe-oop): 0.051 Å |
| O | chainID: C, resSeq: 402, molecule: OXYGEN ATOM |
|||
9qqy-E-401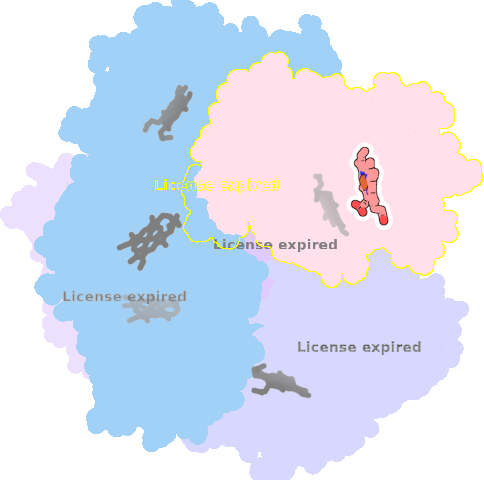 |
sad. +0.35 ruf. +0.36 dom. +0.07 bre. -0.07 |
HIS | chainID: E, resSeq: 225, molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup |
oxidoreductase oligomeric count: 6 pocket vol.: 434.0 Å3 d(Fe-oop): 0.032 Å |
| O | chainID: E, resSeq: 402, molecule: OXYGEN ATOM |
|||
9qqy-F-401 |
sad. +0.30 ruf. +0.48 dom. -0.04 bre. -0.22 |
HIS | chainID: F, resSeq: 225, molecule: Putative dye-decolorizing peroxidase (DyP), encapsulated subgroup |
oxidoreductase oligomeric count: 6 pocket vol.: 442.0 Å3 d(Fe-oop): 0.094 Å |
| Not assigned. | ||||
