Browser
Protein data
function: oxidoreductaseexperiment: X-RAY DIFFRACTION
resolution: 1.93 Å
origomeric count: 26
axial ligand #1: HIS
chainID: N,resSeq: 61,
Coordination distance[Å]: 2.026
molecule: Cytochrome c oxidase subunit 1
EC number: 1.9.3.1
Organism: Bos taurus
axial ligand #2: HIS
chainID: N,resSeq: 378,
Coordination distance[Å]: 2.08
molecule: same as ligand #1
List of other hemes in pdb:7thu
| ID of heme | Distortion | Axial ligands on heme | Function & structure | |
|---|---|---|---|---|
7thu-A-606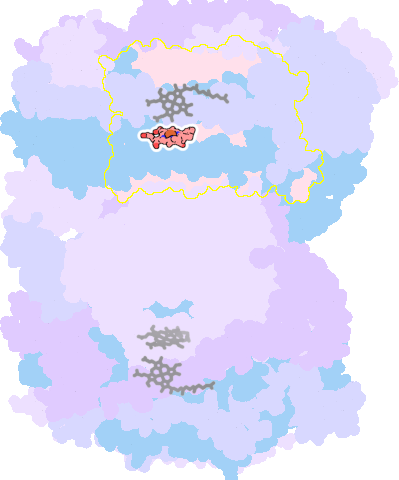 |
sad. -0.45 ruf. -0.00 dom. +0.58 bre. -0.31 |
HIS | chainID: A, resSeq: 376, molecule: Cytochrome c oxidase subunit 1 |
oxidoreductase oligomeric count: 26 pocket vol.: 697.0 Å3 d(Fe-oop): 0.275 Å |
| Not assigned. | ||||
7thu-A-607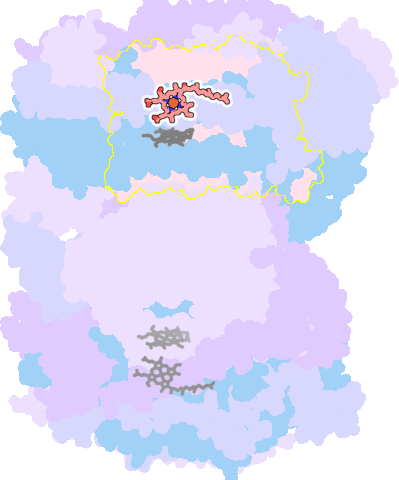 |
sad. -0.69 ruf. +0.40 dom. +0.19 bre. -0.33 |
HIS | chainID: A, resSeq: 61, molecule: Cytochrome c oxidase subunit 1 |
oxidoreductase oligomeric count: 26 pocket vol.: 606.0 Å3 d(Fe-oop): 0.010 Å |
| HIS | chainID: A, resSeq: 378, molecule: Cytochrome c oxidase subunit 1 |
|||
7thu-N-606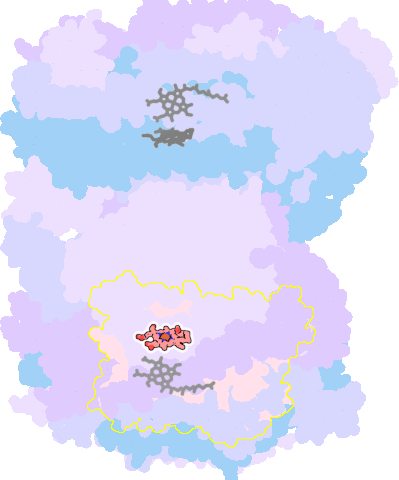 |
sad. -0.42 ruf. +0.00 dom. +0.55 bre. -0.36 |
HIS | chainID: N, resSeq: 376, molecule: Cytochrome c oxidase subunit 1 |
oxidoreductase oligomeric count: 26 pocket vol.: 707.0 Å3 d(Fe-oop): 0.264 Å |
| Not assigned. | ||||
