Browser
Protein data
function: translocaseexperiment: ELECTRON MICROSCOPY
resolution: 2.6 Å
origomeric count: 21
axial ligand #1: HIS
chainID: O,resSeq: 41,
Coordination distance[Å]: 1.923
molecule: Cytochrome c1, heme protein, mitochondrial
EC number: 7.1.1.8
Organism: Mus musculus
axial ligand #2: MET
chainID: O,resSeq: 160,
Coordination distance[Å]: 2.211
molecule: same as ligand #1
List of other hemes in pdb:7o3h
| ID of heme | Distortion | Axial ligands on heme | Function & structure | |
|---|---|---|---|---|
7o3h-C-401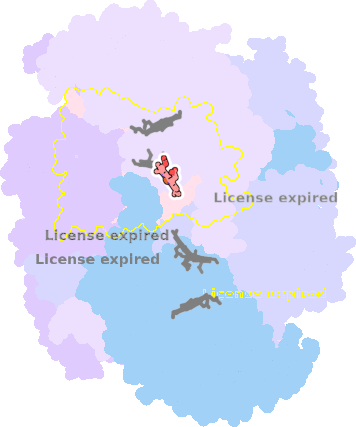 |
sad. -0.00 ruf. +0.36 dom. -0.02 bre. +0.01 |
HIS | chainID: C, resSeq: 83, molecule: Cytochrome b |
electron transporter oligomeric count: 21 pocket vol.: 449.0 Å3 d(Fe-oop): 0.008 Å |
| HIS | chainID: C, resSeq: 182, molecule: Cytochrome b |
|||
7o3h-C-402 |
sad. -0.13 ruf. -0.88 dom. +0.16 bre. +0.08 |
HIS | chainID: C, resSeq: 97, molecule: Cytochrome b |
electron transporter oligomeric count: 21 pocket vol.: 411.0 Å3 d(Fe-oop): 0.055 Å |
| HIS | chainID: C, resSeq: 196, molecule: Cytochrome b |
|||
7o3h-D-301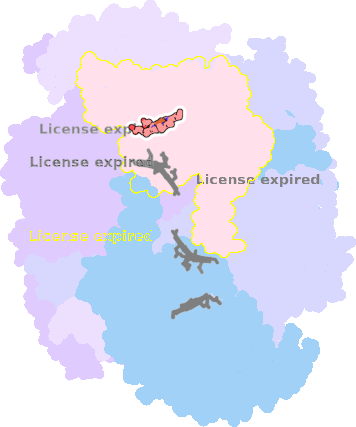 |
sad. +0.05 ruf. -0.03 dom. -0.08 bre. -0.19 |
HIS | chainID: D, resSeq: 41, molecule: Cytochrome c1, heme protein, mitochondrial |
translocase oligomeric count: 21 pocket vol.: 436.0 Å3 d(Fe-oop): 0.054 Å |
| MET | chainID: D, resSeq: 160, molecule: Cytochrome c1, heme protein, mitochondrial |
|||
7o3h-N-401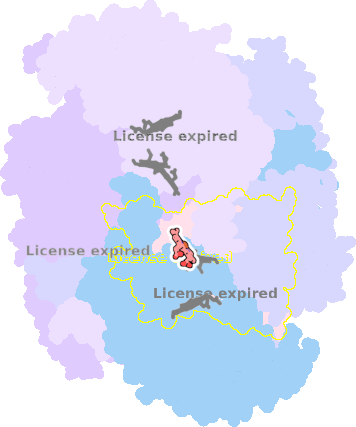 |
sad. -0.01 ruf. +0.44 dom. +0.04 bre. +0.02 |
HIS | chainID: N, resSeq: 83, molecule: Cytochrome b |
electron transporter oligomeric count: 21 pocket vol.: 443.0 Å3 d(Fe-oop): 0.015 Å |
| HIS | chainID: N, resSeq: 182, molecule: Cytochrome b |
|||
7o3h-N-402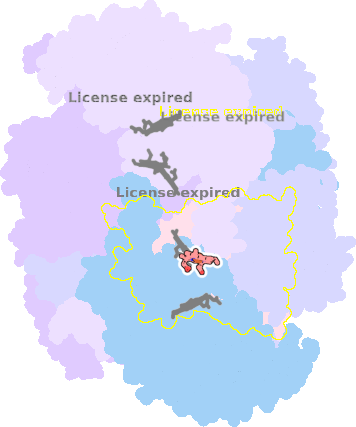 |
sad. -0.13 ruf. -0.99 dom. +0.14 bre. +0.09 |
HIS | chainID: N, resSeq: 97, molecule: Cytochrome b |
electron transporter oligomeric count: 21 pocket vol.: 422.0 Å3 d(Fe-oop): 0.050 Å |
| HIS | chainID: N, resSeq: 196, molecule: Cytochrome b |
|||
